Trade-offs in the sizes of visual and olfactory organs are a common feature of animal evolution, but the underlying genetic and developmental mechanisms have not been clear. A study publishing August 22 in the journal Development Cell reveals that a single DNA variant that affects the timing of sensory organ development in fruit flies could explain the size trade-off between eyes and antennae, potentially providing a quick route to behavioral changes and adaptation.
Because the affected gene, eyeless/Pax6, is conserved across invertebrates and vertebrates, including humans, the discovery could represent a general mechanism for sensory organ size trade-offs across the animal kingdom.
The senses animals rely on have been shaped through evolution to better navigate and exploit the environment. As a result, even closely related species living in different ecological niches show variation in the sizes and shapes of their sensory structures. In arthropods such as fruit flies, trade-offs between the size of the eyes and of the antennae, where most olfactory organs are located, are pervasive.
“What we demonstrate is that there are consequences to subtle changes in the conserved mechanisms that govern how these sense organs develop,” says senior study author Bassem Hassan of Institut du Cerveau et de la Moelle épinière (Institut du Cerveau – ICM). “What this means more broadly is that one cannot fully understand how genetic variation and morphological variation relate to each other without understanding the developmental processes that translate the former into the latter.”
To examine the underlying mechanisms, Hassan and first author Ariane Ramaekers of Institut du Cerveau et de la Moelle épinière (Institut du Cerveau – ICM) combined comparative analyses of different fruit fly strains and species with developmental, molecular, and genome-editing approaches. Specifically, the authors focused on a structure called the eye-antennal imaginal disc (EAD), which consists of an eye field and a non-eye field and gives rise to all external head sensory organs during fruit fly development.
Hassan and Ramaekers found that the eye field is proportionally larger in Drosophila pseudoobscura (D. pse.) compared to Drosophila melanogaster (D. mel.), corresponding to a 35% increase in the number of ommatidia—small units that make up the insect compound eye. Similarly, the eye field is proportionally larger in the D. mel. strain called Canton-S compared with the D. mel. strain Hikone-AS, corresponding to a 12.5% increase in ommatidia number.
“These data suggest that, despite 17 to 30 million years of separated evolution between the two species groups, ommatidia number variation between D. mel. and D. pse. and between two D. mel. strains share a common developmental logic,” Hassan says.

Heads from D. melanogaster (top) and D. pseudoobscura (bottom) show different proportions of eyes and antennae. (Image credit: A. Ramaekers and N. Grillenzoni).
To search for genetic causes of eye size variation, the researchers next examined DNA sequences that transcription factors bind to regulate the expression of the neighboring eyeless/Pax6 gene. They found that a single nucleotide variant—a G>A substitution—at a binding site that differentiates the small eye subspecies from the large eye subspecies. The G allele in small eye subspecies is predicted to have a higher affinity for the binding site, resulting in greater repression of eyeless/Pax6 gene expression compared to the A allele in the large eye subspecies.
Additional analyses showed that this variant occurs in natural fruit fly populations, and the A allele corresponds to both more ommatidia and smaller antennal width among different laboratory strains. Using CRISPR/Cas9 to introduce the A allele into a G-homozygous stock, the researchers demonstrated that the G>A substitution causes an increase in the number of ommatidia.
“We were surprised by the simplicity of the mechanism of sensory trade-offs that we identified: To vary sensory organ size—in this case, eye versus antennae—it suffices to slightly vary the expression of a single gene,” Ramaekers says. “It was particularly satisfactory to find that this gene, called Pax6, is the same one that builds the eye in all animals, including humans. We were also surprised that what matters to generate a trade-off is to change when, rather than where, Pax6 ends up being expressed. To make the eye bigger or smaller, it is sufficient to slightly speed up or slow down the subdivision of the head primordium into eye versus non-eye territories.”
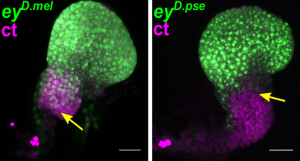
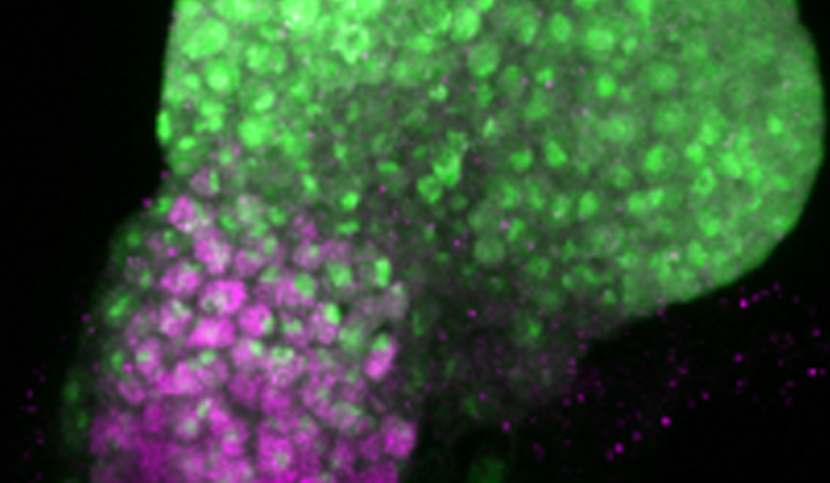
Different temporal regulation of ey/PAX6 from D. melanogaster (left) and D. pseudoobscura (right). During the late second larval stage, the activity of ey/PAX6 enhancer from D. pseudoobscura, visualized by the expression of a reporter gene (in green, GFP), has almost entirely disappeared from the non-eye portion of the EAD (labelled in magenta, ct immunostaining). In contrast, at the same developmental stage, the D.melanogaster enhancer activity is still detected in this compartment. (Image credit: A. Ramaekers).
For the authors, the findings raise several intriguing questions. For example, it’s not yet clear how the single-nucleotide variant changes the timing of Pax6 expression and sensory organ development. Moreover, it’s possible that controlling the timing of the expression of key developmental genes could be a general rule for changing the size of a tissue or organ. Another interesting question is whether sensory brain regions are affected by changes in the relative sizes of sensory organs.
###
This work is supported by VIB, the Belspo WiBrain Interuniversity Attraction Pole network, Fonds Wetenschappelijke Onderzoeks (FWO), the Institut Hospitalier Universitaire (IHU), the Institut du Cerveau et de la Moëlle Epinière (Institut du Cerveau – ICM), the FLiACT Marie Curie ITN, and a Swiss National Science Foundation (SNSF) grant.
Developmental Cell, Ramaekers et al.: “Altering the temporal regulation of one transcription factor drives evolutionary trade-offs between head sensory organs” https://www.cell.com/developmental-cell/fulltext/S1534-5807(19)30658-6DOI: 10.1016/j.devcel.2019.07.027